How I Built an Offline Medical Transcription App and Kept Correcting Myself Along the Way
I'm building an app that listens to a doctor talk to a patient and writes the clinical note for them. The whole thing runs on a laptop. No internet, no cloud, no audio leaving the device.
The idea is simple: a doctor hits record, talks to their patient, and when they're done, a structured clinical note is sitting there ready to review. Two AI models do the work. One hears audio and types out words (like a really fast transcriptionist), and the other reads that transcript and writes the actual clinical note from it.
This is about how I chose those two models, the problems I ran into, and how I kept having to correct my own assumptions along the way.
Voice Picking the Voice Model
This was the easy decision.
The popular choice for voice-to-text is Whisper. It's well-known, supports 99+ languages, and works reasonably well for general speech. But I'm not building a general speech app. I'm building a medical one, and medical speech is a different animal. Doctors say things like "amlodipine" and "bilateral crackles." Whisper hears "am low to peen" and "by lateral crackles."
Google released a model called MedASR through their Health AI Developer Foundations program. MLX support (the framework that lets AI models run on Apple Silicon) landed on March 7th, 2026. A week ago. The numbers made this an easy call:
Word Error Rate
MedASR: ~5% on medical words
Whisper: 25-32%
That's a 5-6x accuracy difference.
Model Size
MedASR: 105M parameters
Whisper: 1.5B (15x larger)
MedASR fits in 8GB RAM.
Training Data
MedASR: 5,000 hours of clinical audio (physician dictations, conversations)
Whisper: general internet speech.
That's not a close call. I went with MedASR.
Pipeline Fixing the Voice Pipeline
Then I hit a bug that taught me something about how these models work. A doctor says "thirty year old male patient." The app transcribes it as "three year old male patient."
The problem: greedy decoding. MedASR processes audio in tiny slices and picks the best-matching word for each slice in isolation, with no memory of previous picks and no awareness of what's coming next. The word "three" sounds almost identical to the beginning of "thirty," but "three" exists as a single token in MedASR's vocabulary while "thirty" has to be assembled from four pieces (th + ir + t + y). By the time the "-ty" arrives, the model has already committed to "three."
The fix: Google ships a 672 MB language model alongside MedASR -a massive dictionary of word patterns that knows which phrases are common in medical English. It was already on my machine. I just wasn't using it. Once plugged in, the model considers 8 possible interpretations simultaneously instead of committing one at a time. The language model knows "thirty year old" is far more probable than "three year old" in a clinical context. I also added hotword boosting: a list of clinical terms and number words that get a 10x weight boost when audio is ambiguous.
Google's benchmarks showed this reduces word errors by about 30%:
| Dataset | Without language model | With language model |
|---|---|---|
| RAD-DICT | 6.6% errors | 4.6% errors |
| GENERAL-DICT | 9.3% errors | 6.9% errors |
| FM-DICT | 8.1% errors | 5.8% errors |
There was a second issue: corrupted audio. MedASR expects clean audio at 16,000 samples per second. The app's browser environment captures at 48,000 and was downsampling badly -just picking every third sample and discarding the rest. High-frequency sounds like the "t" in "thirty" were getting mangled. On top of that, browser features designed for video calls (echo cancellation, noise suppression, auto gain) were processing the audio in ways MedASR wasn't trained on.
The fix: Let the browser handle sample rate conversion natively (much higher quality than our manual approach), disable all video-call audio processing, and add standard cleanup (DC offset removal and dither). Combined with the language model fix, these changes contributed to roughly 30% fewer word errors overall.
Eval The First Eval
Before I ran my own evaluation, there was already one. A teammate had evaluated 7 models using Promptfoo, an open-source evaluation framework, with Gemini 3 Pro as the judge. It was a solid piece of work: 24 synthetic patient transcripts, 2 prompt variants, 7 rubric criteria per output, 336 scored results total.
The results looked great:
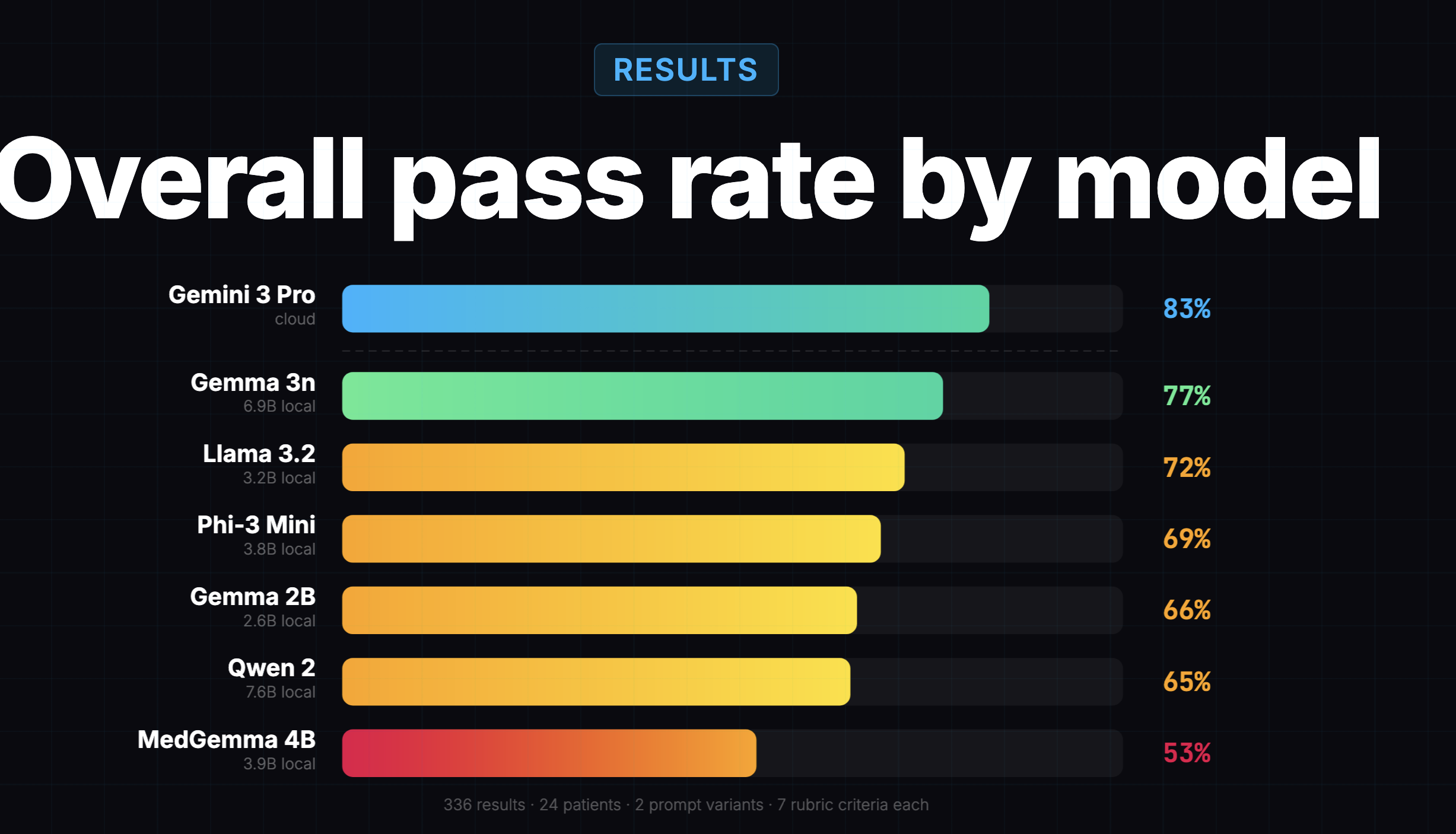
Gemma 3n scored 77%, only 6 points behind the cloud model at 83%. The conclusion was "local models are surprisingly viable," and Gemma 3n was the obvious winner.
That's the eval I used to pick Gemma 3n in the first place. And to be fair, it got a lot right. It correctly identified that entity tagging was hard for all local models. It correctly showed that formatting and clinical quality were easy problems (95%+ across the board). It provided a useful directional signal about which models were worth considering.
Something Felt Off
But as I built the offline app and started reading the notes Gemma 3n actually generated, something felt off. On simple visits, the notes were fine. On complex visits -multi-condition patients with labs, medications, and counseling -the notes were thin. Important details were missing. I'd read the transcript, then read the generated note, and think: where did half this visit go?
I went back to the earlier eval's heatmap and found the clue hiding in plain sight:
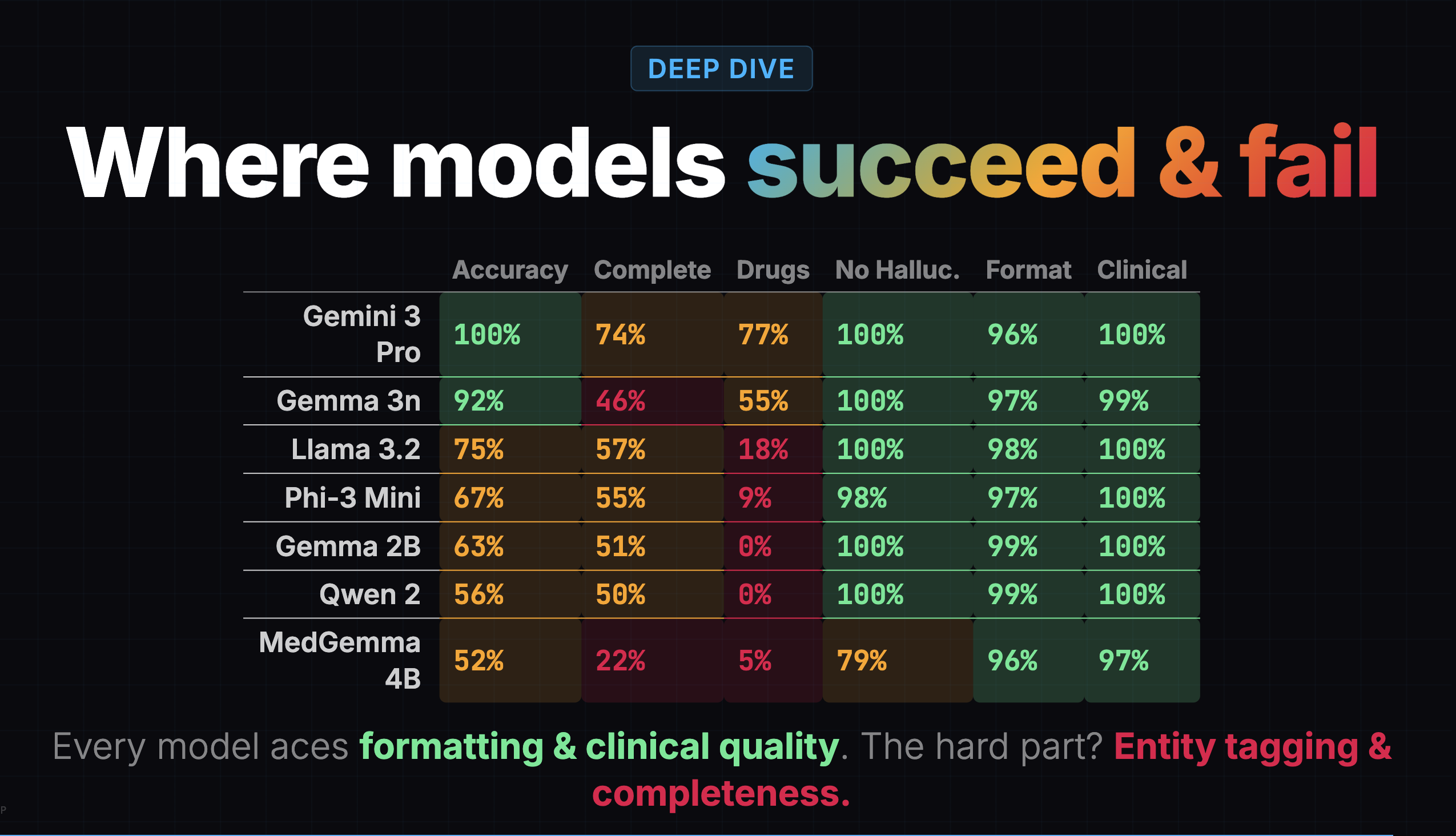

Gemma 3n's completeness score was 46%. It was capturing less than half the clinical content. But the overall score of 77% buried this because formatting (97%), clinical quality (99%), and hallucination resistance (100%) were all near-perfect. The model was producing well-formatted, professional-sounding notes that were missing half the information.
A confident-looking note with missing content is arguably worse than a messy note with everything in it, because the doctor might not realize something is missing.
Rubric Same Model, Different Score
The earlier eval wasn't wrong, exactly. It measured what it set out to measure. But there were things it couldn't tell me that I needed to know.
The most striking example is MedGemma 4B. In the earlier eval: 53% overall, dead last, 22% completeness, 79% hallucination (worst of any model). In my eval: 49% overall, 48% completeness, 53% hallucination. Same model, dramatically different picture. The difference wasn't the model. It was the rubric.
MedGemma has a distinctive style: it writes more than other models. It captures lab values, exam findings, counseling details, and clinical boilerplate that other models skip. But it's messy about it. Content lands in the wrong section. Standard clinical phrases get included without being prompted.
The old eval's rubric was ambiguous on these edge cases, and the scores suggest the judge interpreted that ambiguity harshly. My eval addressed this with explicit instructions: "If content is present but placed in the wrong section, penalize template adherence only. Do NOT also penalize completeness." The hallucination rubric now exempts four standard clinical defaults by name: "alert and oriented," "no known drug allergies," "well-appearing," and "in no acute distress."
MedGemma went from worst to essentially first among local models not because anything changed about the model, but because the scoring system went from ambiguous to explicit on the edge cases that affected it most.
Method Running My Own Eval
Beyond the rubric differences, there were structural issues I wanted to address:
- Binary pass/fail masked severity. My eval used a 0–3 scale, which revealed that the difference between "captures some content" and "captures most content" matters a lot in clinical documentation.
- Single judge, no cross-check. My eval used Claude Opus as the primary judge and Gemini 2.5 Pro as a secondary judge. The two mostly agreed on rankings.
- Equal weighting hid what matters. My eval weighted hallucination and completeness at 2x, entity marking at 0.5x.
I tested 8 models: 7 that run on a laptop, plus one cloud model (Gemini) as a benchmark. The local models ranged from 3 to 4 billion parameters, all 4-bit quantized to fit in 8GB of RAM on an Apple Silicon Mac. (Quantization compresses each parameter into a smaller representation, shrinking a 4B model from 8-16 GB to roughly 2-2.5 GB -some precision lost, like compressing a high-res photo, but it's what makes local inference possible.)
I ran all 8 against 25 doctor dictation transcripts, generating 200 SOAP notes total. Two AI judges scored every note across 6 dimensions: hallucination, completeness, instruction following, template adherence, entity marking, and duplication.
The initial results said Gemma (the model I was already using) was beating MedGemma by 8 points, 49% to 41%. Case closed, right?
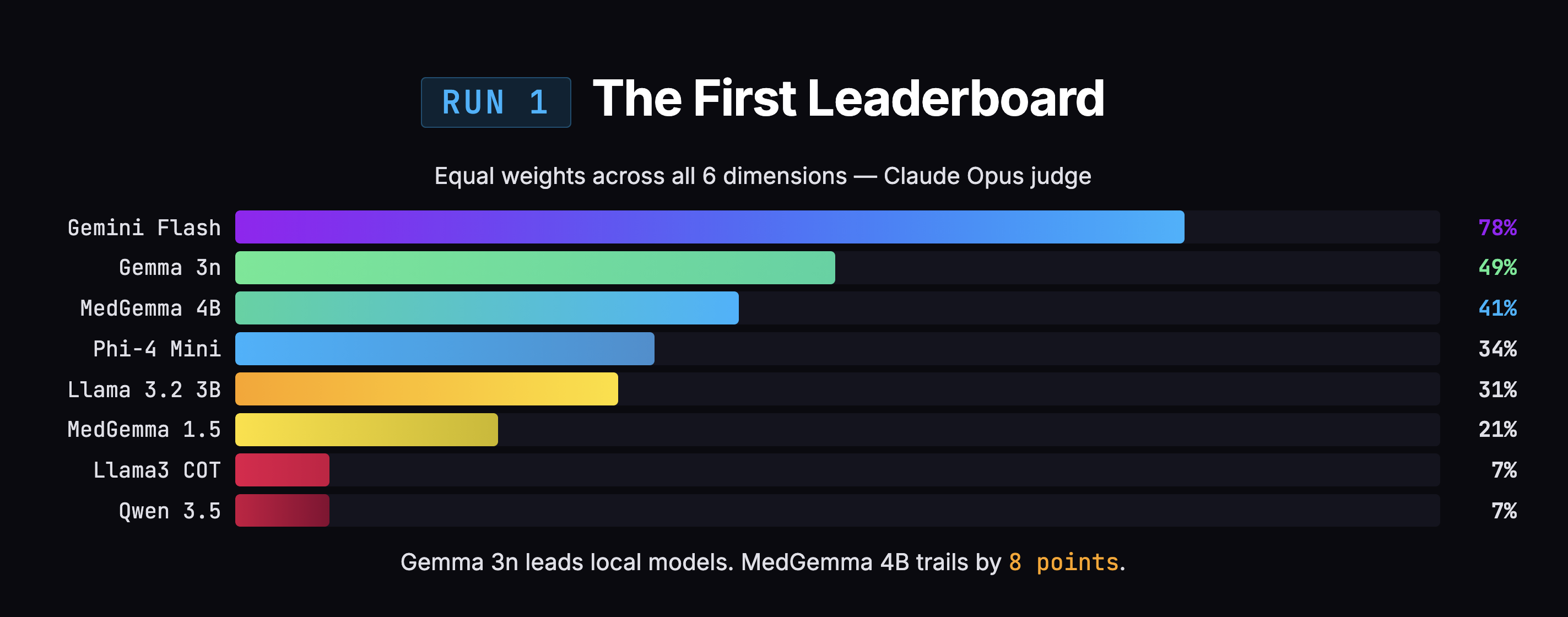
Problems Grading My Own Grading
Not quite. When I looked closer at how the scores were being calculated, I found three problems with my own rubric.
1. Double-penalization. When a model captured the right clinical information but put it in the wrong section, the judge docked points for both "you missed content in the right section" and "you put content in the wrong section." Same information, penalized twice. This disproportionately hurt MedGemma, which captures more information but organizes it messily.
2. Clinical boilerplate flagged as hallucination. Phrases like "alert and oriented" and "no known drug allergies" show up in virtually every real clinical note. But because the doctor didn't explicitly say these words, the judge flagged them as fabricated content. That's like penalizing a note-taker for writing "Meeting started at 2 PM" when nobody announced the time out loud.
3. Equal weighting. Entity marking was broken for every local model (0-3% scores), making it a non-differentiating dimension that still dragged down overall scores. Meanwhile, hallucination and completeness -the dimensions that actually matter for patient safety -carried the same weight as formatting. A doctor can fix bad formatting in two seconds. A doctor cannot fix missing information without re-listening to the recording.
The Scores Flipped
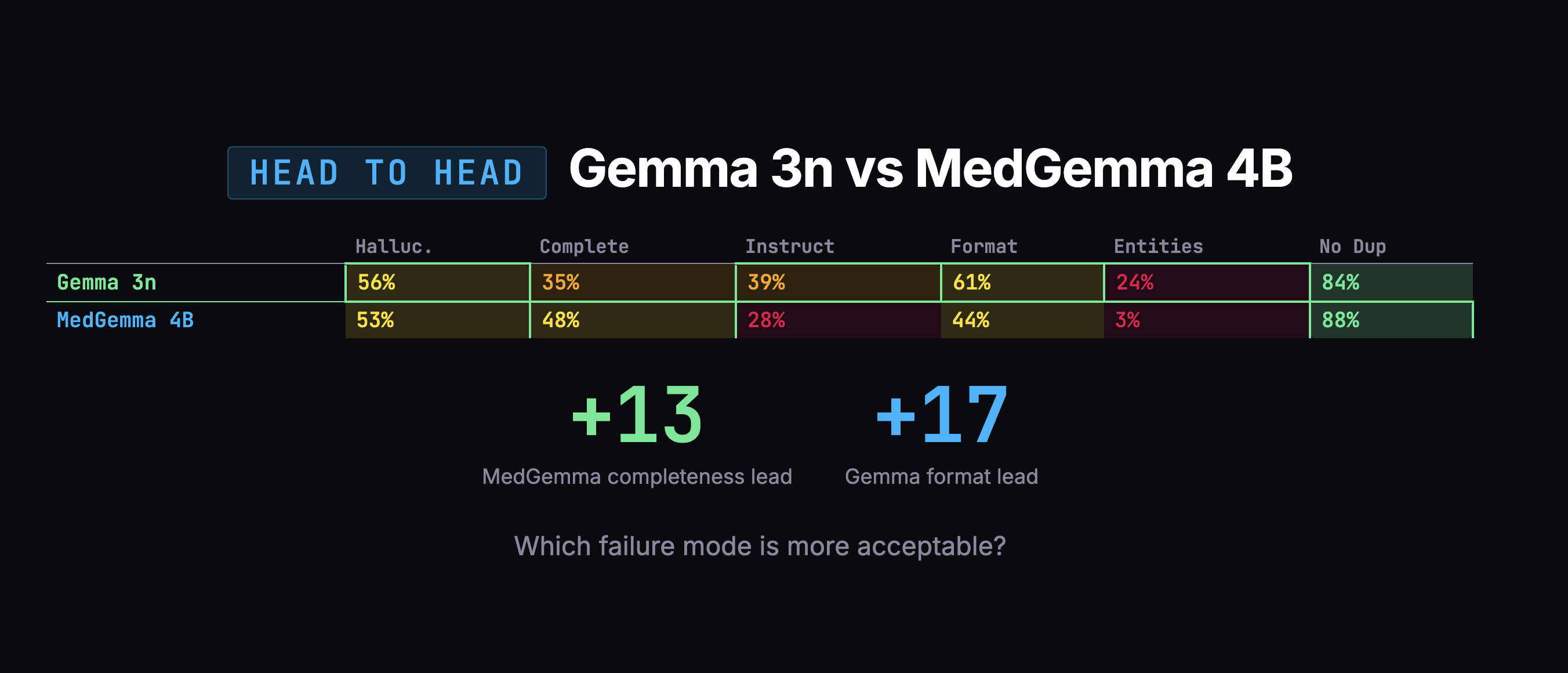
After fixing those three issues and re-scoring all 200 notes with both judges, the 8-point gap between Gemma and MedGemma collapsed to 1 point (50% vs 49%). Statistically meaningless. MedGemma jumped +8 points. Gemma moved +1.
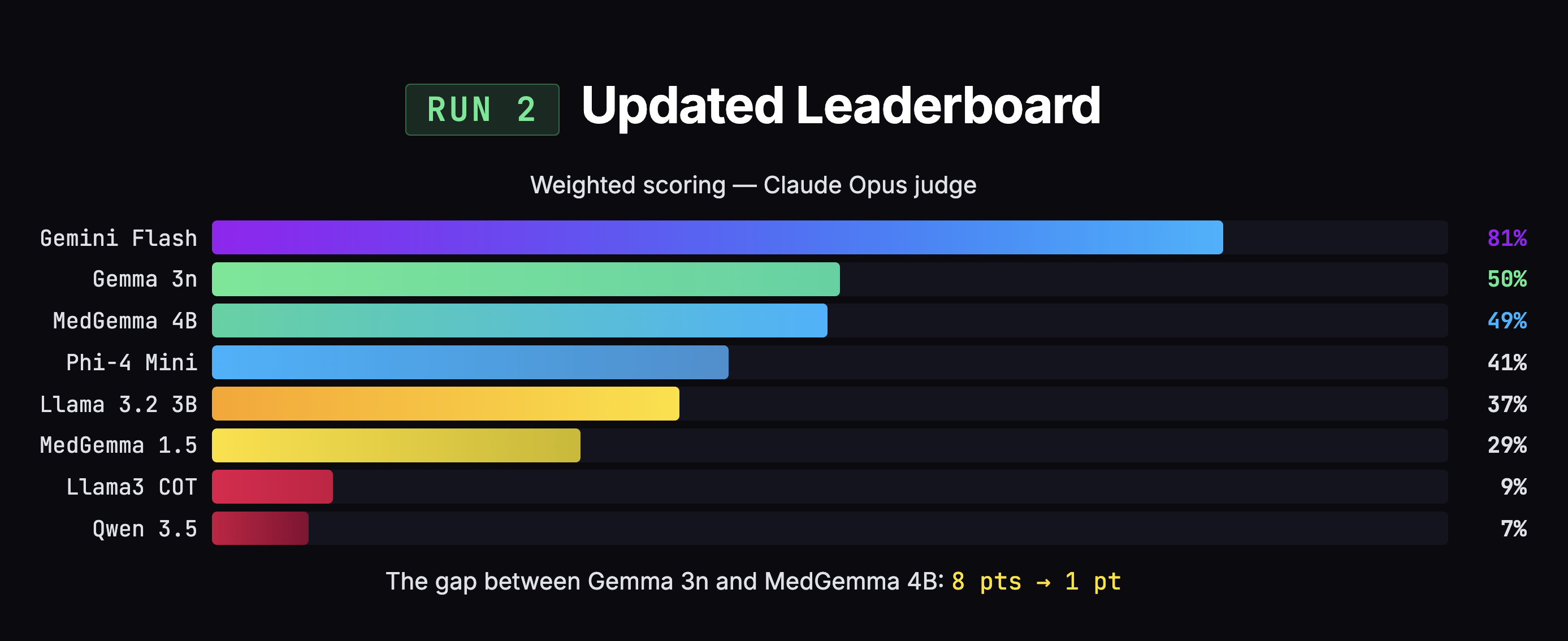
But the scores weren't the whole story. When I looked at what actually matters -how much clinical content each model captured -MedGemma grabbed 48% of the information versus Gemma's 35%. That's a 13-point advantage on the dimension that matters most.

Decision Choosing by Failure Mode
The overall scores were too close to decide on numbers alone. So I looked at how each model fails. Gemma's failures were scarier: confusing which family member had a medical condition, duplicating entire notes. MedGemma's failures were mostly messy organization and verbosity -annoying, but fixable in post-processing.
I swapped engines.
Format Fighting the Model's Training
There was one more thing I had to fix, and it was my own fault.
I'd been telling all these models to format note headings using bold text (**Subjective** instead of # Subjective). Every AI model was trained on text where # means a heading. Asking a small model to use bold text as a heading is like asking someone fluent in English to write in a dialect they've never seen.
MedGemma was scoring only 24% on instruction following, and this formatting mismatch was a major reason why. Once I switched to standard markdown headings, I removed a whole category of unnecessary errors -and more importantly, removed what would have been a tax on every future fine-tuning run.
Next Fine-Tuning: Closing the Gap
There's a 32-point gap. But here's the critical insight from the evals: MedGemma's failures aren't about medical knowledge. It understands medicine. It just doesn't know my specific output format: my entity tagging, my section structure, my expected level of detail. These are behavioral gaps, not knowledge gaps. And that's exactly what fine-tuning is designed to fix.
The plan: distillation via LoRA. Use Gemini as a teacher. Show it 1,700 real doctor-patient conversations, collect the notes it writes, run them through a two-pass quality filter, and train MedGemma to mimic that output across 7 note templates. LoRA freezes the original model and trains a small adapter on top (1-10M new parameters) -like putting a contact lens on the model rather than performing surgery. The adapter is small (10-100 MB), so training is fast and cheap.
The target is 65-75% quality (up from 49%), roughly closing the gap to Gemini by half or more. Total cost: about $10-15 in API calls. Training time: hours on a Mac.
Future What's Next
The plan is to distill from Gemini 3 Pro and re-run evals with four models side by side: Gemini 3 Pro (the new ceiling), Gemini 2.5 Flash (the old ceiling), vanilla MedGemma 4B (the baseline), and our fine-tuned MedGemma (the result). That comparison will tell us exactly how much of the gap we closed.
And here's why doing evals first mattered: the training examples will use the corrected markdown format, the corrected scoring rubric, and they'll target the specific model I selected because of the evals. Without evaluating first, I might have spent days fine-tuning the wrong model with the wrong format, only to discover later that I needed to start over.
The Thread
Looking back, the pattern is obvious. I kept correcting myself:
- Corrected the voice pipeline (plugged in the language model that was already sitting there)
- Corrected the evaluation rubric (three flaws found and fixed)
- Corrected the model choice (swapped from Gemma to MedGemma)
- Corrected the formatting approach (stopped fighting model pre-training)
- Corrected the approach order (evals before fine-tuning, because you can't fine-tune a model you haven't chosen yet)
None of these corrections came from planning better upfront. They came from looking at what was actually happening and being willing to change direction. The evals didn't just help me pick a model. They revealed that my scoring was flawed, my formatting was fighting the models, and my assumptions about which model was better were wrong.
If I had just picked a model and started fine-tuning without evaluating first, I'd be polishing the wrong thing right now.